Repeat Expansions - Long and Short Read Sequencing
I want to try and do a series of posts comparing long and short reads for a few different applications. This isn’t intended as a deep drive exactly, but a bit of a learning process for me and an interesting exercise.
The first thing I’m looking at is repeat expansions. There are various reasons for this, not in the least because I’ve found myself reading about one particularly common repeat expansion (FMR1). But also because it should be an application where we see a clear difference between short and long reads.
First… what is FMR1?
FMR1

FMR1 is a gene on the X chromosome. This gene contains a trinucleotide (CCG) repeat. Normally this repeat is ~40 repeat units long. Polymerases and other enzymes don’t generally like repeats. As such there’s a tendency to slip in this region… and the repeat expands.
If it expands to ~200 units it causes a large scale instability on the X chromosome called fragile X. You can see this on Karyotypes:
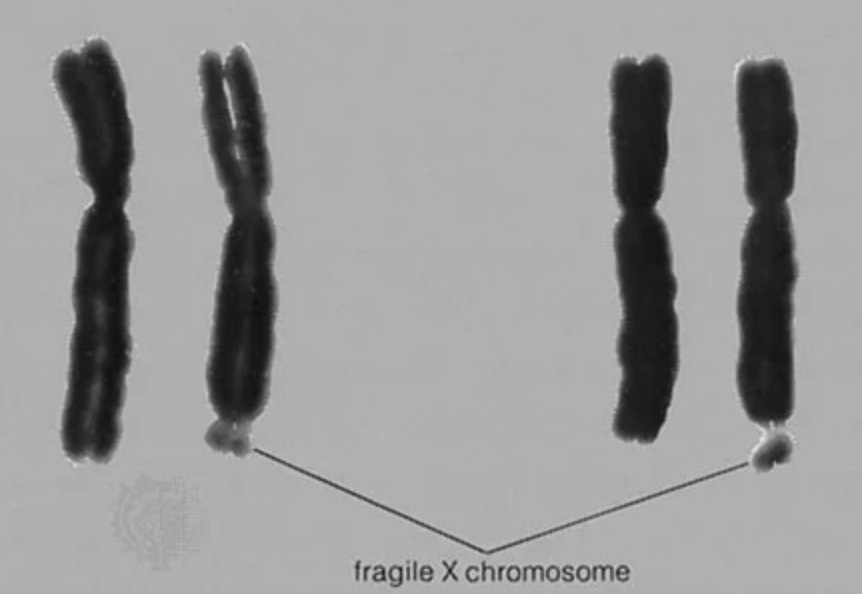
It is also a leading monogenic cause of intellectual disability. What is called “fragile X syndrome”.
Sequencing
Fragile X isn’t normally tested for via sequencing. But this and other repeat expansions could be tested for as part of a whole genome sequencing based diagnostic. In particular because repeats are… long. Long read sequencing should have an advantage here.
In this evaluation I’ll be using HG002 . That’s because datasets from Illumina, PacBio and Oxford Nanopore exist here making a direct comparison easy.
Pileups
First let’s just look at the data.
Illumina
Here’s the FMR1 repeat expansion region in a Illumina pileup:
This pileup is kind of confusing (at least to me). Reads spanning this region start ok, but as they get into the repeat region most reads show deletions before matching on the other side. It’s clear that there’s something going on… but I don’t have much confidence in my ability to interpret the data from the pileup alone…
PacBio
If we now look at a long read (PacBio) pileup of the same region things are much clearer, we have a 33bp insert with respect to the reference:
Eyeballing this suggests the PacBio data shows 30 copies of the repeat unit.
Oxford Nanopore
Oxford Nanopore data seems far more confusing:
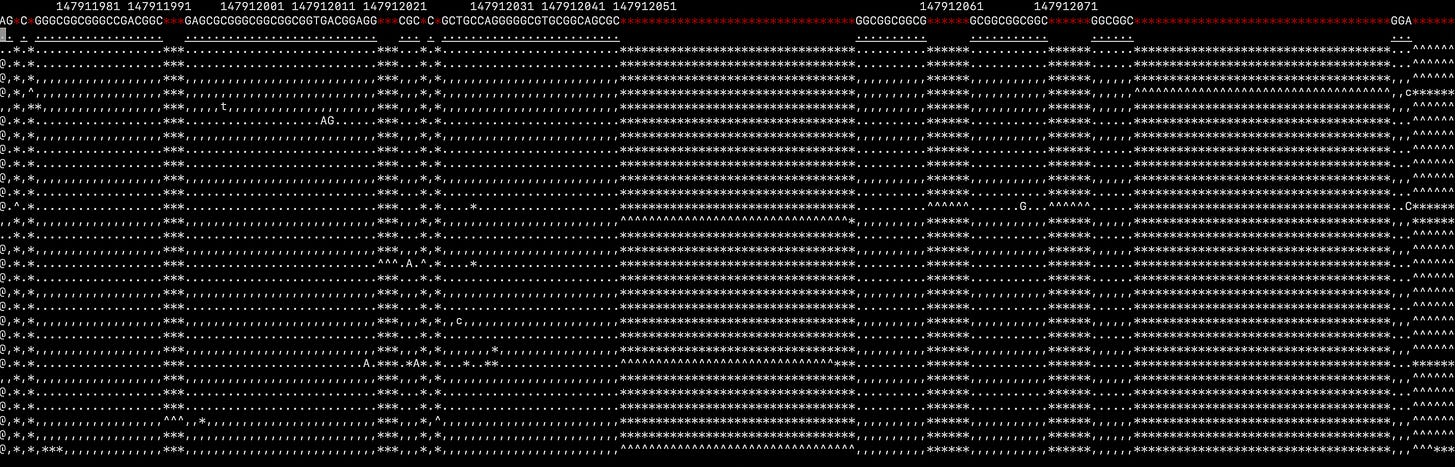
Zooming out it seems like the alignment may have been throw off somehow…
Oxford Nanopore’s platform seems to have significant issues with repeat regions and this is probably what we’re seeing here. Some algorithms might help correct for these issues.. but it seems harder to interpret than the short read data.
Illumina - ExpansionHunter
To make sense of the Illumina data we need to use algorithms. In the appendices below I explain how I installed (appendix A) and ran (appendix B) ExpansionHunter. In appendix C I discuss the complexity of this tool. The fact that there are 3 appendices might suggest this is a non-trivial process… It should also be noted that a PCR-free prep appears to be required here (I assume to avoid the potential from PCR bias).
In any case it generated a genotype call of 31.
Conclusion
PacBio data seems quite nice here. You can clearly see the repeat expansion without using any fancy algorithms. However, short read data can also provide a reasonable estimate of the repeat expansion length.
For clinical applications, I suspect the slightly better estimates you get from PacBio data may not be clinically relevant. However I’m still convinced that it’s the better dataset and cost no issue you can make a strong case for a PacBio genome being “the clinical standard”.
I’d like to follow this up by looking at datasets containing the full mutation (200 repeat units). Others have shown impressive work using long reads here.
However this whole exercise took far too long. And more subscribers showing interest here would help motivate this.
What I’m saying is please subscribe.
I’d also welcome comments and suggestions via email (new@sgenomics.org or Discord). This is a first pass on an unfamiliar datatype, so it’s entirely possible there are errors.
Appendix A - Installing ExpansionHunter
ExpansionHunter is on GitHub here. The installation instructions aren’t in the top level README.md and when found it’s a multi-step process. So for me this fails the Joel Test. But the builds starts relatively easily, so I’m only moderately irritated.
It then spends a few minutes compiling seemingly every known C++ library before crashing with:
/mnt/sdd1/family/analysis/ExpansionHunter/build/googletest-prefix/src/googletest/googletest/src/gtest-death-test.cc:1301:24: error: ‘dummy’ may be used uninitialized [-Werror=maybe-uninitialized]
1301 | StackLowerThanAddress(&dummy, &result); | ~~~~~~~~~~~~~~~~~~~~~^~~~~~~~~~~~~~~~~So I just hack the code such that “dummy=0” at initialization and go on with my day… well that is to say I wait another few minutes… until it crashes again with:
/mnt/sdd1/family/analysis/ExpansionHunter/ehunter/thirdparty/graph-tools-master-0cd9399/src/graphcore/GraphCoordinates.cpp:197:54: error: ‘::max’ has not been declared; did you mean ‘std::max’?
197 | uint64_t result = std::numeric_limits<uint64_t>::max();Probably this is something from a new C++ which isn’t in my relatively recent version of g++ on my relatively recent Ubuntu distribution… I just give it good old UINT64_MAX. I have to make similar changes in 2 other places…
Wait… more… minutes…
Yay 1/1 Tests passed and binaries! Well I assume there are binaries somewhere among the other 92842 files…
If you wrote ExpansionHunter… I’m sorry. It’s not your fault.
Appendix B - Using ExpansionHunter
To use ExpansionHunter we need a variant catalog file. We want to look at FMR1 so we go searching for this and find a random comment on biostars. Which contains this variant catalog information.
We don’t know which human reference this is for.. so we’re going to have to figure that out I guess.
{
"FMR1_CGG": {
"ReferenceRegion": "chrX:147911919-147951125",
"VariantType": "Repeat",
"TargetRegion": "chrX:147911919-147951125",
"RepeatUnit": "CGG"
}
}In any case we find out at this point that our alignment BAM wasn’t indexed so we have to wait for that to run…
And then try to run ExpansionHunter which results in a different error:
ExpansionHunter --reads ./XXX.bam --reference ./GCA_000001405.15_GRCh38_no_alt_analysis_set.fna.gz --variant-catalog ./FMR1.json --output-prefix exprun --threads 24
2023-12-10T08:11:00,[Starting ExpansionHunter v5.0.0]
2023-12-10T08:11:00,[./FMR1.json is not a path to an existing file]FMR1.json absolutely exists. Trying a full path doesn’t work… Googling the error doesn’t find anything… so we run strace and see this:
newfstatat(AT_FDCWD, "./FMR1.json\302\240", 0x7ffef1e4acb0, 0) = -1 ENOENT (No such file or directory)It looks like we’ve picked up some weird whitespace characters on the command line, so we put some new whitespaces in and rerun which fails with:
2023-12-10T08:17:50,[Starting ExpansionHunter v5.0.0]
2023-12-10T08:17:50,[Analyzing sample Natsuki_NG1VFVR45X]
2023-12-10T08:17:50,[Initializing reference ./GCA_000001405.15_GRCh38_no_alt_analysis_set.fna.gz]
[E::fai_build3_core] Cannot index files compressed with gzip, please use bgzip
Segmentation fault (core dumped)The Segfault after the error is just the icing on the cake. But it’s kind of clear we need to decompress the reference.
Now it runs. But then fails with:
2023-12-10T08:23:14,[Starting ExpansionHunter v5.0.0]
2023-12-10T08:23:14,[Analyzing sample Natsuki_NG1VFVR45X]
2023-12-10T08:23:14,[Initializing reference ./GCA_000001405.15_GRCh38_no_alt_analysis_set.fna]
2023-12-10T08:23:27,[Loading variant catalog from disk ./FMR1.json]
2023-12-10T08:23:27,[Field LocusId must be present in {"FMR1_CGG":{"ReferenceRegion":"chrX:147911919-147951125","RepeatUnit":"CGG","TargetRegion":"chrX:147911919-147951125","VariantType":"Repeat"}}]Looking at the VariantCatalog documentation it seems like the format has changed since whatever random nonsense is present in the Biostars comment. So let’s try the example in the documentation:
[
{
"LocusId": "FMR1",
"LocusStructure": "(CGG)*",
"ReferenceRegion": "X:146993568-146993628",
"VariantType": "RareRepeat",
"OfftargetRegions": [
"12:7781290-7781350",
"12:125052154-125052156",
"16:25703613-25703635",
"16:28074516-28074518",
"17:30814024-30814026",
"17:64298467-64298469",
"19:2015524-2015526",
"2:87141540-87141618",
"2:92230909-92230911",
"2:211036020-211036032",
"2:225449878-225449880",
"20:30865500-30865516",
"5:443334-443364",
"7:20824939-20824941",
"7:100271437-100271439",
"7:104654597-104654599",
"7:143059853-143059855",
"9:100616695-100616697",
"X:20009036-20009046"
]
}
]The reference locations are completely different… which is nice. But the software does run:
2023-12-10T08:28:55,[Starting ExpansionHunter v5.0.0]
2023-12-10T08:28:55,[Analyzing sample XXXX]
2023-12-10T08:28:55,[Initializing reference ./GCA_000001405.15_GRCh38_no_alt_analysis_set.fna]
2023-12-10T08:28:55,[Loading variant catalog from disk ./FMR1.json]
2023-12-10T08:28:55,[Running sample analysis in seeking mode]
2023-12-10T08:28:56,[Analyzing FMR1]
2023-12-10T08:28:57,[Writing output to disk]But the results don’t look right… randomly googling reference positions above it seems these are specified against hg19. I’m using hg38 (GCA_000001405.15_GRCh38_no_alt_analysis_set.fna).
At least some random webpage gives us the FMR1 positions in hg38… but what about all the off target regions? What are they for anyway?
Anyway I guess we just throw them into the first online tool we find and get:
[
{
"LocusId": "FMR1",
"LocusStructure": "(CGG)*",
"ReferenceRegion": "X:147912050-147912110",
"VariantType": "RareRepeat",
"OfftargetRegions": [
"12:7628694-7628754",
"12:124567608-124567610",
"16:25692292-25692314",
"16:28063195-28063197",
"17:32487006-32487008",
"17:66302349-66302351",
"19:2015525-2015527",
"2:86914417-86914495",
"2:92042883-92042885",
"2:210171296-210171308",
"2:224585161-224585163",
"20:32277697-32277713",
"5:443219-443249",
"7:20785320-20785322",
"7:100673814-100673816",
"7:105014150-105014152",
"7:143362760-143362762",
"9:97854413-97854415",
"X:19990918-19990928"
]
}The JSON output then gives us a Genotype of “31” confidence interval 31-31.
I’m sure that’s fine!
Appendix C - ExpansionHunter Complexity
Post build the ExpansionHunter directory contains 92842 files. Just counting the cpp files this is 1.7M (1749493) lines of code and 0.3M lines in headers. For reference a truly massive project like Google Chrome contains ~6M lines of code. Is Google Chrome really only 3x more complicated that ExpansionHunter.
ExpansionHunter performs a single, relatively simple analysis. The paper doesn’t go into a huge amount of detail, but it seems to be performing a graph based realignment of reads around repeat expansions using regular expressions as the input definition:
I’m going to be honest. I suspect this code be implemented in significantly less than 10000 lines of code, using a custom implementation. Google Chrome in its 6M line code base has 60000 open issues. A total of 1.5M appear to have been filed across the course of the project. ExpansionHunter would have 500K issues if it accumulated bugs at the same rate or 20000 if we just look at open issues.
This is a long way of saying that there are probably bugs in ExpansionHunter because of bug surface is so large. A lot of these files are probably not used by ExpansionHunter directly… but it’s still a concern.
More importantly ExpansionHunter has potential clinical applications. And this large bug surface is a concern there too.
Of course it’s not just ExpansionHunter. In this pipeline we’ve had to use Samtools and Minimap. Tools which have been designed largely with research applications in mind.
For clinical applications. We might want to consider better tooling.




