PumpkinSeed
GenomeWeb published an article about 3 months after mine on PumpkinSeed. Which is fine… it’s not a competition.
Overall I think my previous article was pretty good! A few new pieces of information have appeared.
Two applications targets:
deSIPHR technology sequences peptides up to 30 amino acids — potentially more
cell-MAPP (cell monitoring across phenotype progression) assays the ability of peptides to activate immune cells.
Collaborations:
Genentech to identify immunopeptides for oncology
IMEC for fab on 300mm wafers, able to manufacture chips with around 100 million sensors per cm squared.
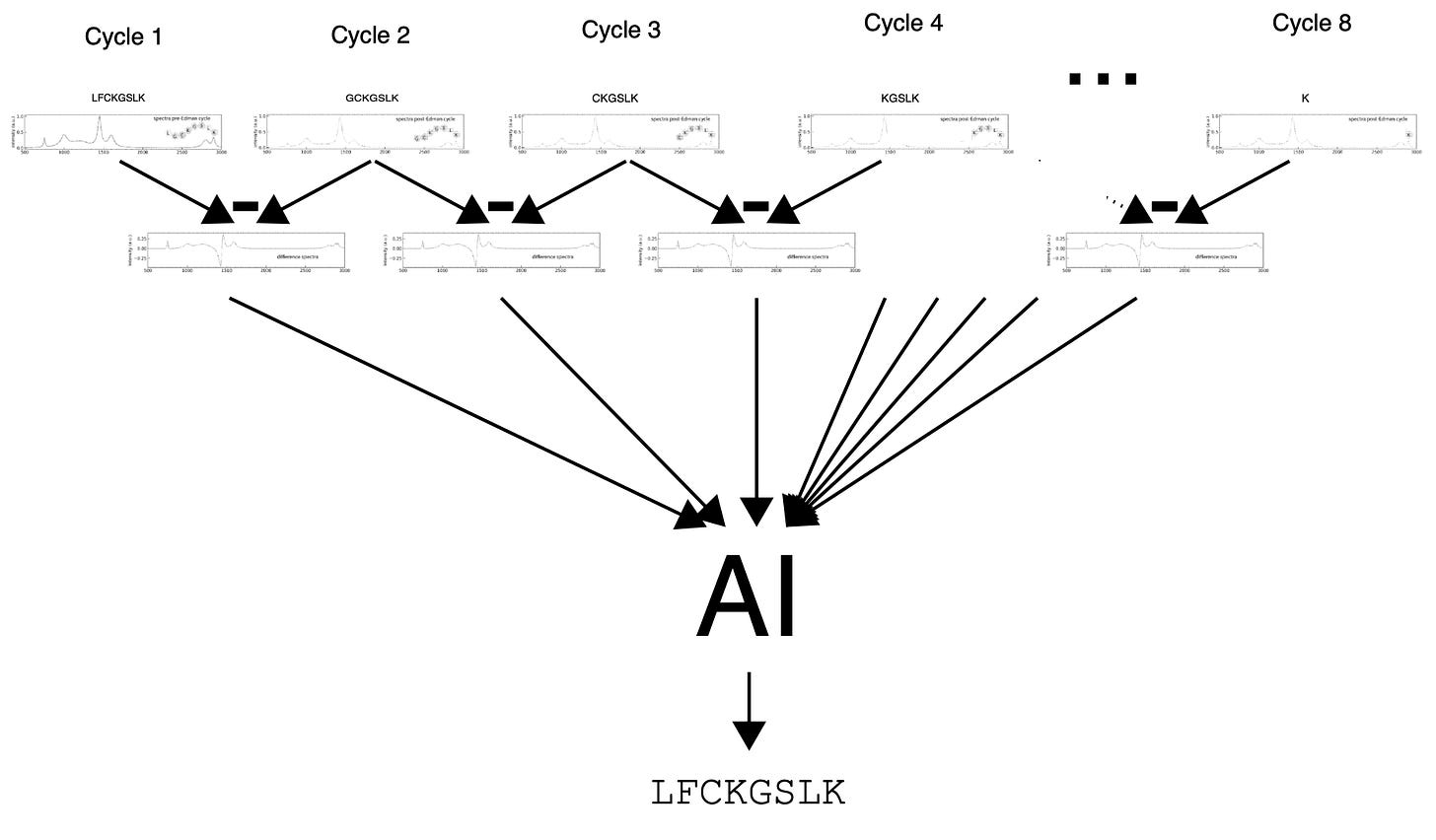
For sequencing, they said this:
“For peptides, the sequencing is done by subtraction. After initial spectral fingerprints for a molecule are obtained, the terminal ends are cleaved enzymatically, and new spectra are obtained at each step.”
Which I think is more or less what I said.
A couple of more interesting quotes below!1 but essentially confirming the single molecule Raman, nano-antenna nature of the platform.
Interesting stuff!
“The silicon is enhancing light, boosting Raman scattering by over eight orders of magnitude, allowing us to detect single molecules using label-free methods,” Dionne said.
“All nucleated cells have protein fragments displayed on their cell surface that essentially tell your immune system whether a cell is healthy or diseased,” Dionne said. “Figuring out what those kind of protein fragment sequences or peptide sequences are is pretty challenging, but once you have figured out those sequences, which is what deSIPHR does, you want to know if those sequences actually activate a particular set of patient T-cell responses.” Cell-MAPP offers that functional information, she said.

A couple of questions:
1) How do they plan to capture a single peptide per feature?
2) How many features could they realistically have per chip? For proteins presumably you'd like a high number of peptides due to the dynamic range..